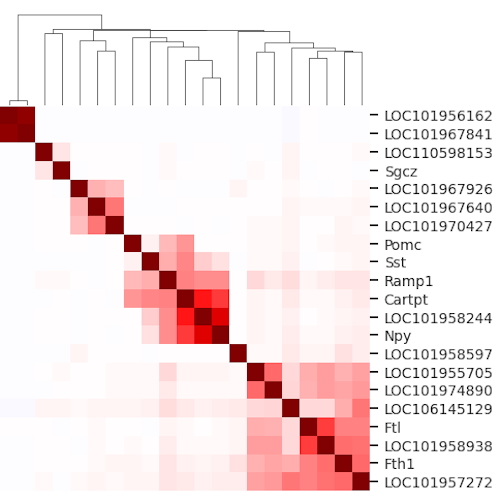
Bioinformatics
Developing methods for challenging datasets
In brief: I develop robust pipelines for sequencing data.
Industry bioinformatics strives to convert raw data to medically actionable insights. To achieve this, pipelines need to be simple, robust, and attributable. In my industry work at Fauna Bio, I balanced dual objectives: on one hand, keeping up with the state of the art in single-cell analysis; on the other, ensuring the analysis logic is transparent enough to trace procedures and their assumptions from the raw reads to a proposed drug target.
I designed and adapted a diversity of methods to answer questions about the biology of hibernation. Using unique internal single-nucleus RNA sequencing datasets, I constructed statistically robust analyses to identify cell types, improve differential expression testing, detect patterns of cell–cell signaling, infer gene regulatory networks, and study rhythmicity in hibernator gene expression. To apply these insights to human health, I closely collaborated with my colleagues to integrate them into the company’s drug discovery platform, as well as to drastically speed up existing analysis workflows.
To facilitate analysis and ensure reproducibility, I developed and deployed automated analysis pipelines in cloud environments. To improve the broader bioinformatic infrastructure, I have taken the opportunity to contribute to open-source projects, such as Nextflow.
Single-cell sequencing produces vast amounts of actionable data, yet interpreting it requires care and understanding of its chemical and biological context: common artifacts can distort or mislead interpretation. I have deeply explored the pitfalls of sequencing analyses, and characterized previously unreported isolation and sequencing artifacts.
Non-model organism bioinformatics presents unique challenges: the genome references are often incomplete. I have a substantial experience base in annotating de novo references, as well as using public resources to diagnose and account for errors in existing references and data.